Blogs & News
Using Hadoop for Lung Volume Calculation from Computed Tomography
This post outlines an initial investigation into distributed image analysis with Hadoop. The investigation focuses on an imaginary case study – the calculation of lung volume from a CT scan Thorax. This is the traditional method of imaging the lung tissues. The technologies and API’s explored are detailed in figure 1. The Python wrap of ITK (included in red) represents a viable option which wasn’t explored as part of this investigation.
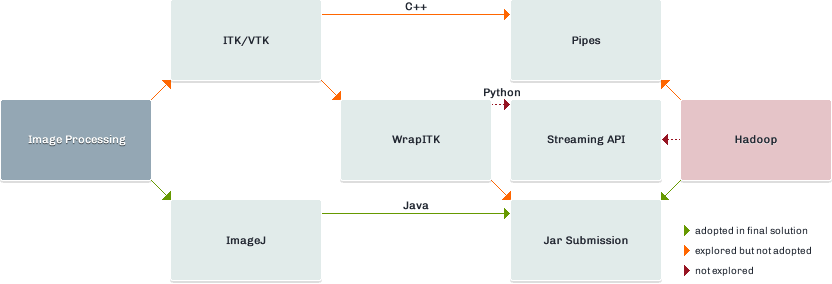
Image Data and Hadoop
- Patient: male
- Area: Thorax/abdomen
- Modality: Computed tomography
- Number of slices: 225
- Physical dimensions (mm): 335 x 335 x 281.25
- Voxel dimensions (voxels): 512 x 512 x 225
- Total size: 113MB
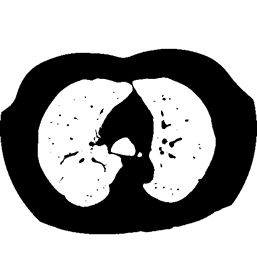
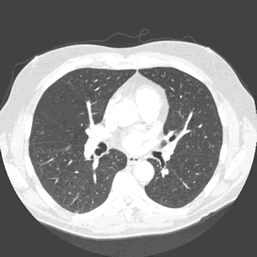
The data used for the case study is detailed on the right. Full size CT scans are in the region of 4GB – 6GB. A sample slice from the three dimensional volume can be seen in figure 3.
SequenceFiles
Due to the relatively small file size of individual slices in the CT scan all image data was written to a SequenceFile prior to staging. This key/value pair format is splittable, supports compression and works with any data which can be serialized to binary. This format allows us to utilise Hadoop’s ability to optimise the staging, distribution and execution of each job through the use of a SequenceFileInputFormat.
Mapper
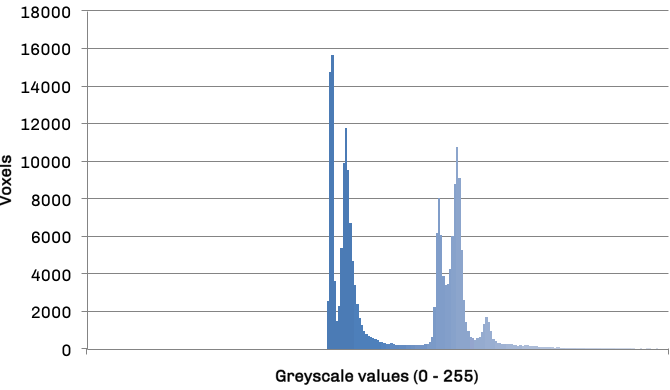
The mapper is responsible for obtaining an area for both left and right lung in each slice, leaving the volume calculations to the reducer. The histogram in figure 3 details the frequency distribution of voxels (volumetric pixels) against their greyscale values for the raw CT slice in figure 2. The two groupings in the distribution represent lower density non-body voxels (including lungs) and higher density tissue voxels respectively. The contrast between tissue and air allows us to effectively apply optimal thresholding to create a binary mask. The result of this process can be seen in diagram figure 4a.
Frequency distribution of voxel values
Vessels within the lungs can be seen as dark speckles in the thresholded image. As these vessels must contribute to the lung volume we apply a binary hole-filling algorithm. The outcome of this process is shown in figure 4b.
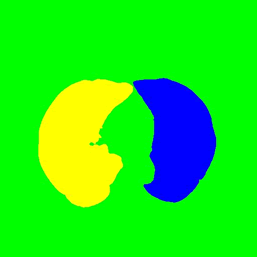
Applying a seeded region growing algorithm to the resultant filled image now allows us to capture the individual areas representing the left and right lung. This process excludes any remaining artefacts (e.g. bronchial tree) from the segmented image. Automation of the seed point selection for the algorithm would be a topic for future investigation. For the purposes of this case study the seed data was passed to the mapper as part of the binary key in the BinarySequenceFile.
The pixel area of right and left lungs in each slice can now be easily identified using the LUT generated by the region growing algorithm and are added to the mapper output under corresponding keys.
Reducer
A reducer will be responsible for each key and associated values; i.e. “left-lung” or “right-lung” and an iterator containing associated areas generated by the mapper. By iterating over these areas, it now becomes trivial to sum them, convert to a volume in voxels and then further convert this volume to its physical representation in litres.
Results
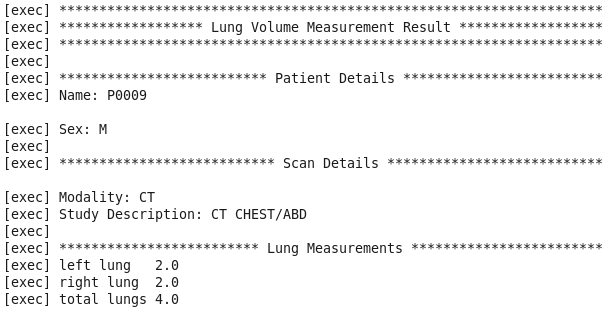
The results from the execution of the MapReduce job can be seen in figure 5. Previous studies have compared this value to that produced by gold standard spirometry. We do not have the spirometry data available for the subject but the value falls within the range expected for a male of his age. Any calculation with the inclusion of spirometry is effort dependent and therefore dependent on breath taken by the patient.
This technique may aid calculation of surgical resection and proportional effect on lung function via volumes. It will not calculate, or make adjustment for, emphysema burden/lung parenchymal disease effect.

March 22, 2013
Harry Peaker
Harry started working at Aridhia in 2013 after graduating with a Bsc(Hons) in Mathematics from the University of Edinburgh. He completed a final year dissertation studying advanced topics in algebra, combinatorics and graph theory, using R and Maple for creating data visualisations and LaTeX for creating reports.
Since joining Aridhia Harry has been involved in a project analysing the human genome – first analysing the output obtained from high-throughput sequencing, and then using APIs to access clinical databases to find up to date clinical relevance for the results.



