Blogs & News
Running AI Agents on Sensitive Research Data: What Is Possible Today
What research teams can do inside a TRE
Most Trusted Research Environments have no AI capability. Researchers who want to use AI tools to work with sensitive data face a straightforward choice: go without or move data to an external service where governance and data residency are someone else’s problem.
Aridhia’s approach, through our AIRA AI framework, is to run AI inference inside the DRE rather than outside it. AIRA runs LLM inference entirely within the DRE’s secure environment using an OpenAI-compatible API, so prompts, outputs, and data derivatives stay on the platform. Administrators control which models are available, every inference job is logged in the audit system, and the same access controls that govern data access apply to AI interactions
This post covers two complementary open-source agentic AI systems we have deployed and tested inside the DRE: Biomni and ClawBio. The tools run on top of AIRA, so they connect to AIRA’s local inference endpoint rather than any external AI service. We use Alzheimer’s disease, rare disease, and cancer as example domains throughout, because they are the kinds of data-intensive research programmes where capable agentic AI has genuine practical value, and where the question of where data goes during analysis cannot be set aside.
Two complementary approaches
Aridhia’s DRE can run almost any agentic AI system that can be containerised. The two tools featured here were chosen because they represent different but complementary approaches to AI-assisted research, and because both are available and working today.
Biomni is a biomedical AI agent developed at Stanford, designed for multi-modal biological reasoning. It approaches a research question the way a knowledgeable collaborator might, forming a plan, deciding what analyses to run, interpreting results, and iterating as findings emerge. It works across heterogeneous data types without needing to be given a fixed sequence of steps, which makes it well suited to open-ended exploratory questions, whether that is reasoning across proteomic and clinical data in an Alzheimer’s cohort, identifying candidate targets in a cancer dataset, or contextualising findings from a rare disease study against the published literature.
ClawBio is an open-source bioinformatics agent skill library integrated with our Aira AI framework inside the DRE. Where Biomni reasons freely across a problem, ClawBio executes specifically defined reproducible workflows. Each skill encodes the correct tools, parameters, and interpretation logic for a specific bioinformatics task, and the agent executes the skill rather than improvising or hallucinating it, which matters considerably when outputs have clinical or regulatory significance.
Local data and reference files, controlled external lookups
Both Biomni and ClawBio run models and core analytical computation entirely within the DRE. Biomni also draws on a substantial library of biomedical reference data stored locally within the secure environment. For many tasks this is sufficient, and no data leaves the environment at any point.
Some workflows do involve outbound reference lookups, however. Biomni performs well when a curated set of public biomedical endpoints is whitelisted, covering literature search, variant databases, pathway resources, and clinical trial registries. ClawBio skills that handle variant annotation and clinical classification similarly query public reference databases for current annotations on variants not held in local databases. In both cases these are read-only lookups to the same public resources a researcher would query manually. No cohort data, no patient records, and no analytical inputs are transmitted outbound. The difference from a researcher doing this work manually is that the lookups happen within a reproducible, auditable workflow running inside the DRE, with egress controlled and logged at the network level.
Aridhia manages this through per-workspace egress whitelisting. The external endpoints permitted are subject to the same audit trail as all other activity within the environment.
What this looks like in practice
The following three examples illustrate the kind of work these tools support today, using Alzheimer’s and rare disease research as the context.
Identifying validated plasma biomarkers for Alzheimer’s disease: Biomni
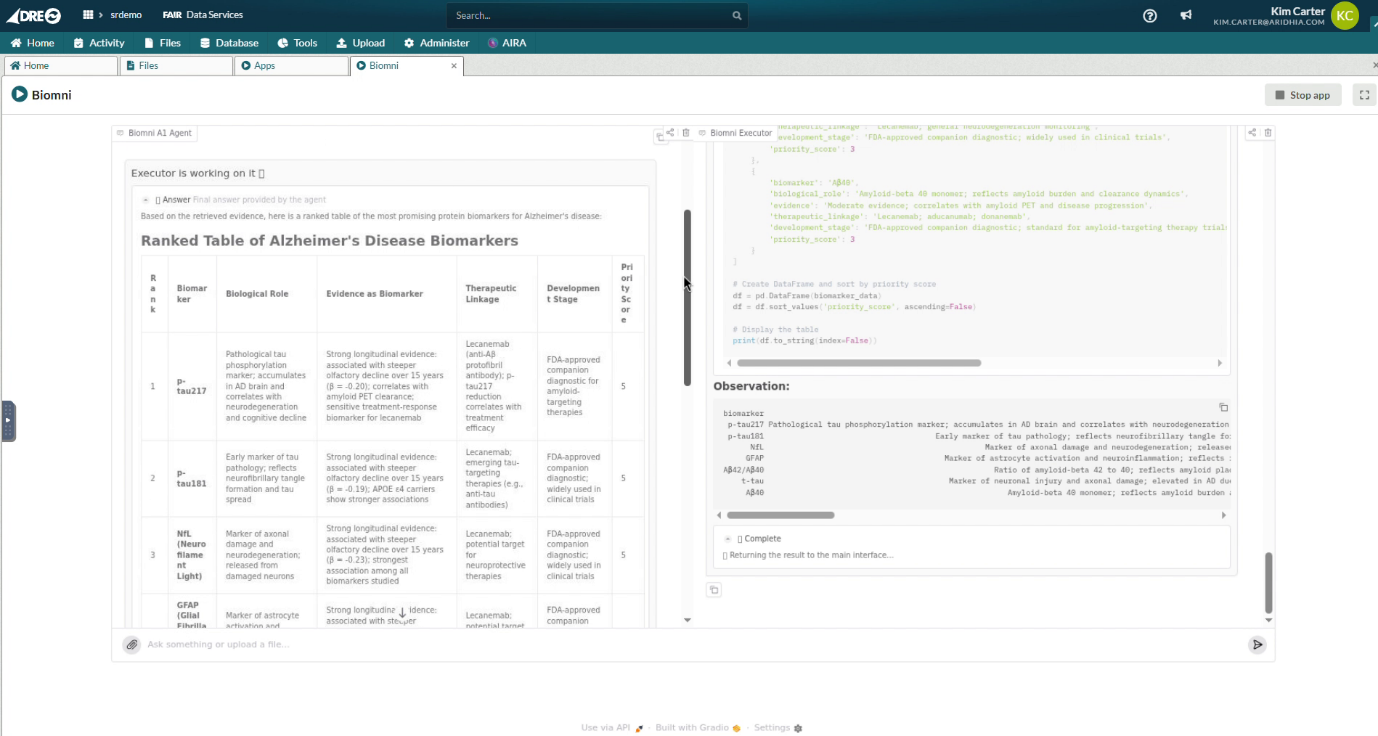
A researcher brings a multi-omics Alzheimer’s cohort into the DRE covering proteomic profiles, longitudinal clinical data, and genetic risk information. Biomni reasons across all of it, ranking candidate protein biomarkers against the evidence base for FDA-cleared plasma assays including p-tau217, GFAP, and NfL, and surfacing where the cohort data converges with or diverges from the validated signal. Where literature context is needed, PubMed is available via controlled egress, but the cohort data does not move. The agent works iteratively, revising its analysis as findings emerge, without the researcher needing to manually orchestrate each analytical step.
[Screen Recording: Biomni session inside the DRE, working through a multi-step reasoning process to produce a ranked table of protein biomarkers for Alzheimer’s disease against user-specified query]
The example shows the ranked biomarker table with supporting rationale from Biomni, in response to the natural language query from the user.
Clinical variant classification: ClawBio ACMG skill
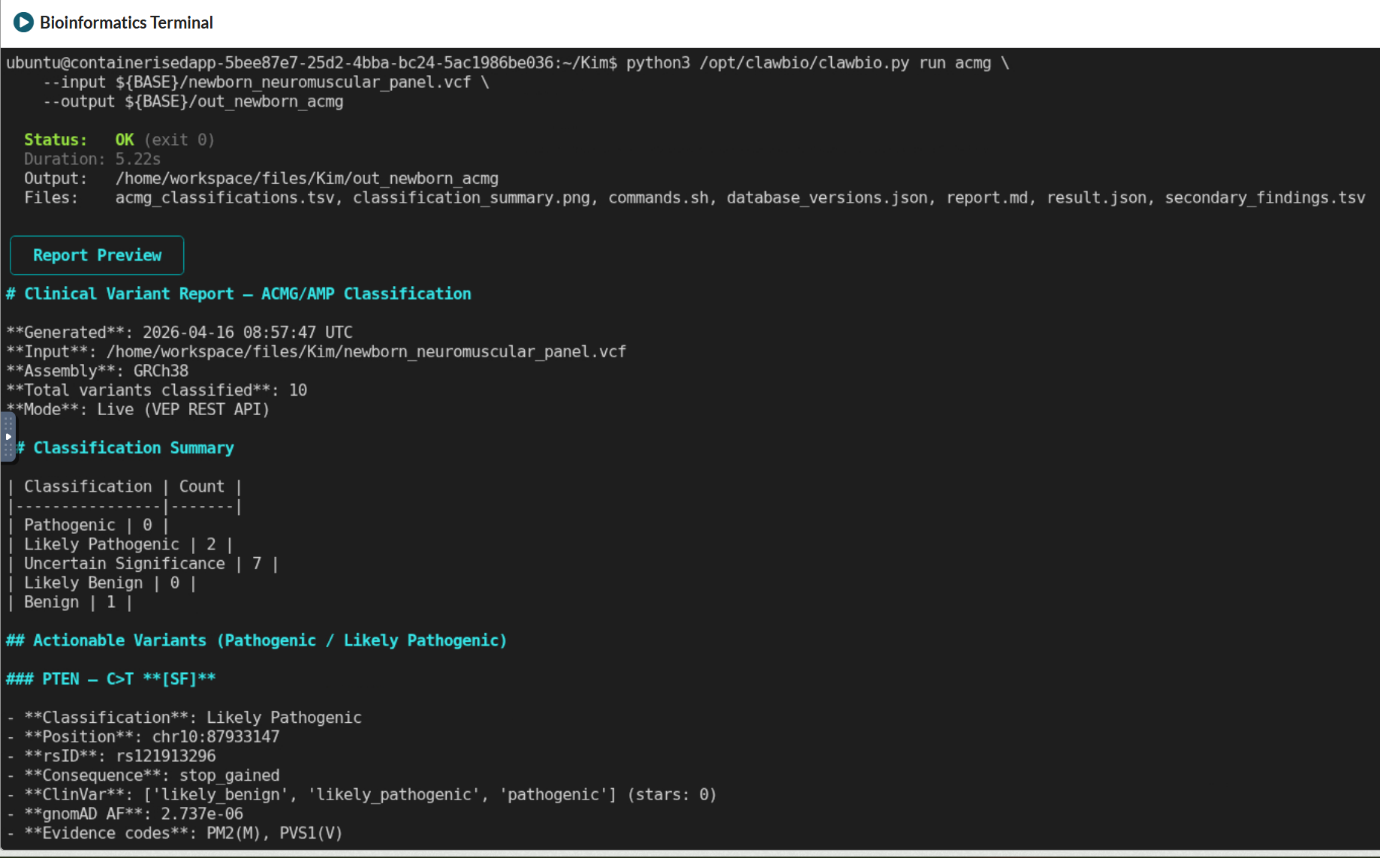
A clinical genomics team working on a rare disease cohort needs to classify variants from a VCF against ACMG/AMP criteria. ClawBio’s ACMG skill runs a 28-criterion classification workflow covering pathogenicity scoring and ACMG Secondary Findings v3.2 screening, entirely inside the TRE. The skill encodes the correct classification logic rather than asking the language model to improvise it, which matters for a workflow where the decisions have direct clinical implications. Output is a structured variant report that is reproducible, auditable, and ready for downstream clinical review.
[Screenshot: ACMG skill running directly via Python against a VCF file in the workspace, showing the 28-criterion classification output in the terminal]
[Screenshot: Claude Code connected to AIRA, receiving a natural language instruction to run the ACMG skill against a workspace file and executing it without data leaving the environment]
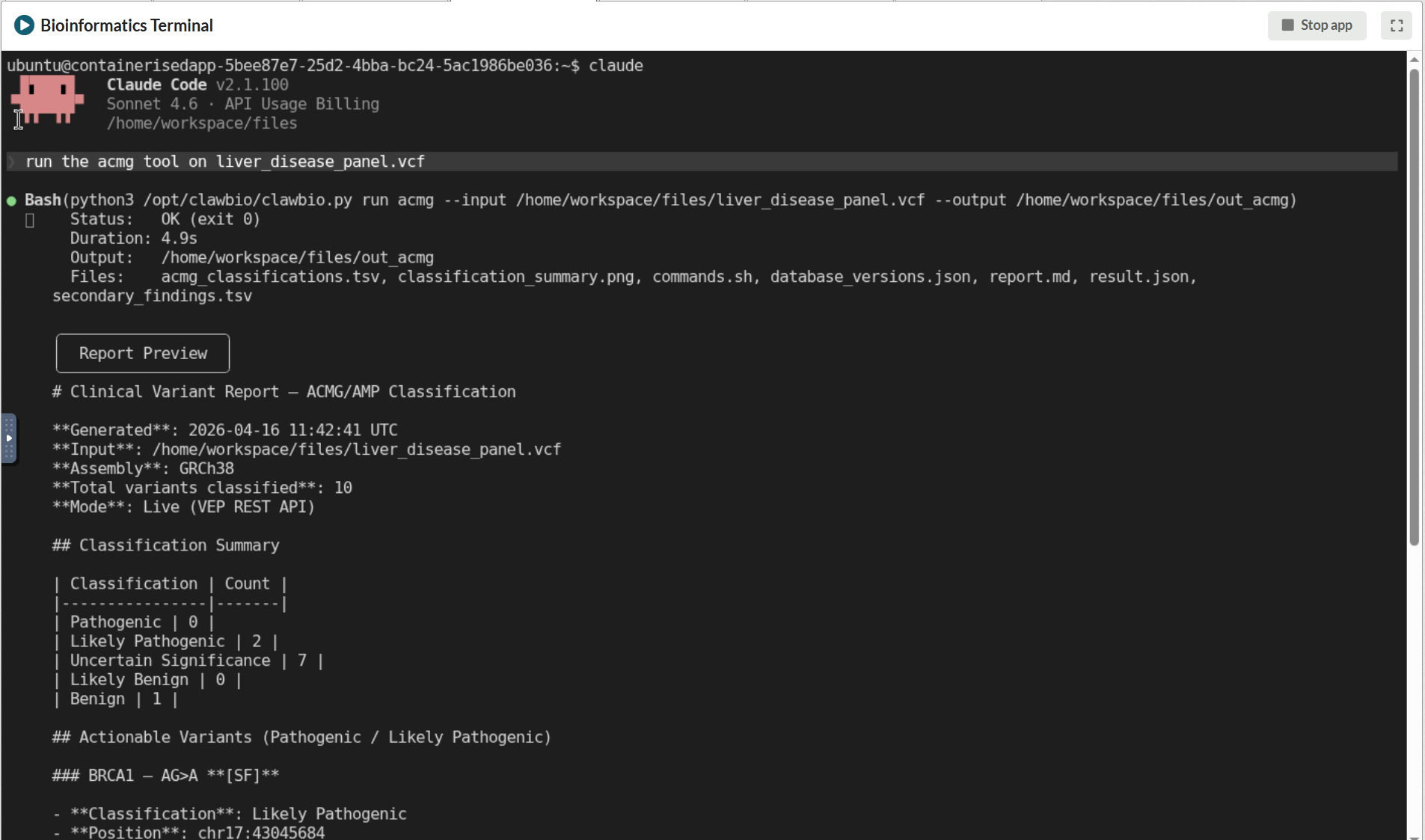
The second example here shows the same skill invoked through Claude Code connected to AIRA, where a natural language instruction is enough to run the full classification workflow against a file already in the workspace. The model handling that instruction is running locally on AIRA, not on any external service.
Polygenic risk scoring: ClawBio PRS skill
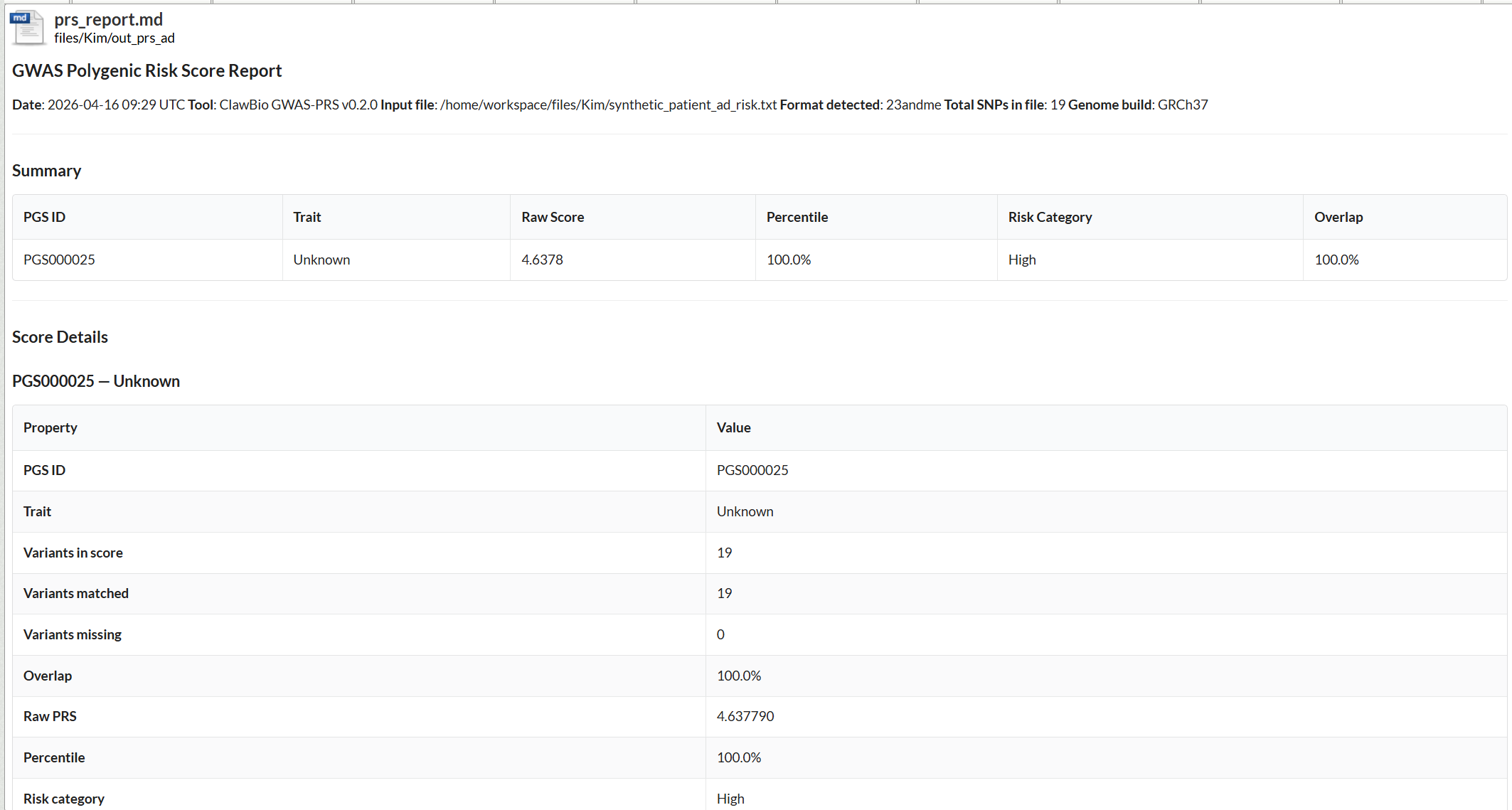
Polygenic risk scores aggregate the cumulative effect of many genetic variants associated with a trait, giving researchers a quantitative measure of individual genetic predisposition. PRS calculated against validated published scores allows patient cohorts to be stratified by genetic risk, supporting case-control comparisons and helping contextualise findings from multi-omics analyses. ClawBio’s PRS skill integrates with the PGS Catalog, which holds over 3,000 published scores across hundreds of traits and diseases. Inside the TRE, a researcher can run PRS on a patient cohort against a published disease score, such as for Alzheimer’s disease, with the skill handling variant matching, score calculation, and percentile ranking against the reference population. The output is a structured report covering matched variants, raw scores, and risk percentiles, produced entirely within the DRE against data that never leaves the environment.
[Screenshot: PRS markdown report output for a synthetic Alzheimer’s disease dataset, showing variant matching, raw score, and risk percentile against PGS000025]
Biomni and agent skills: two approaches to the same problem
Both Biomni and agent skills frameworks like ClawBio, GPTomics bioSkills, and K-Dense AI Scientific Agent Skills put AI to work on biomedical research tasks, but they do so differently, and understanding the distinction helps in choosing the right tool for a given problem.
Biomni is a reasoning agent. It approaches a research question by forming a plan, deciding what analyses to run, interpreting results, and iterating, drawing on broad biomedical knowledge to work across data types and research domains without needing a fixed workflow. The strength is flexibility, generalisability and depth of reasoning across complex, open-ended questions. The trade-off is that like any reasoning system, it can occasionally take an unexpected path, and the analytical decisions it makes are not always fully transparent or reproducible.
Agent skills frameworks work differently. Each skill is a precisely defined, pre-documented workflow with the correct tools, parameters, and interpretation logic written down explicitly and injected into the agent’s context. The agent executes the skill rather than improvising it. The strength is reliability and reproducibility: a variant classification skill applies the same ACMG criteria on every run, a differential expression skill uses a consistent statistical approach, and the outputs are auditable. The trade-off is that skills work best for well-defined tasks and are less suited to open-ended exploratory reasoning where the path is not known in advance.
In practice the two approaches complement each other well. Biomni suits the exploratory phase of a research question, reasoning across a multi-omics dataset, surfacing hypotheses, and identifying where to focus. Agent skills suit the execution phase, running a validated workflow on a specific input and producing an auditable result. A research team with access to both can use each where it is most appropriate, inside the same secure environment and against the same data.
An expanding ecosystem of agent skills
Biomni and ClawBio cover a broad range of biomedical and bioinformatics workflows, and the open-source agent skills ecosystem supporting them is growing rapidly. Aridhia has also tested two further skill libraries inside the DRE: K-Dense AI’s Scientific Agent Skills and GPTomics bioSkills, both of which extend the available analytical capabilities considerably.
K-Dense AI’s Scientific Agent Skills brings 130+ skills spanning domains beyond ClawBio’s current scope, including molecular docking and virtual screening, structural biology, metabolomics, medical imaging workflows, whole slide image analysis, and computational pathology.
GPTomics bioSkills includes more than 400 skills across 60 tightly defined bioinformatics categories, with particular depth on sequencing workflows and strong coverage in areas that neither Biomni nor ClawBio currently address, including CRISPR screen analysis, immunoinformatics and MHC binding prediction. Its spatial transcriptomics skills cover Visium, Xenium, and Slide-seq workflows, and its epidemiological genomics skills support pathogen typing and transmission network analysis relevant to infectious disease and outbreak research.
All four tools run inside the Aridhia DRE as containerised applications under the same governance model, and a research programme is under no obligation to choose just one. A team could use Biomni for multi-modal biomarker reasoning, ClawBio for variant classification, and K-Dense AI’s structural biology skills to assess protein-level impact, all within a single secure environment and against data that never moves.
Staying at the leading edge, without compromise
The agent ecosystem is moving fast, with new tools and model capabilities emerging on a timeline that outpaces traditional software procurement and testing cycles. Aridhia’s approach is to evaluate these tools continuously, testing them inside sandboxed DRE workspaces under secure-environment conditions. Our customers can likewise do this, and gain access to leading-edge agentic AI capability without bearing the integration risk themselves, and without any compromise on the security and sovereignty of their data.
If you are working on a biomedical AI tool or skills library and want to explore what deployment inside a TRE looks like, we would be glad to hear from you. If you are a researcher or research platform team looking for access to the capabilities described in this post, get in touch. We are always looking to bring the right tools to the right data.
We will be posting regularly on this topic as we test and evaluate new tools, sharing what works, what proves difficult, and what it takes to run current agentic AI safely inside a secure research environment.
April 16, 2026
Kim Carter
Dr. Kim Carter is the Chief Data Officer at Aridhia, driving data strategy and empowering customers to maximize the potential of Aridhia's SaaS/PaaS platform. Before Aridhia, Kim was the Senior Manager of Data and Insights at Minderoo Foundation, where he led the global federated cancer data platform and a large D&I team.
Previously, Kim was a Senior Managing Consultant at Data Analysis Australia. He also served as Manager of Bioinformatics at the Telethon Kids Institute (10 years), where he developed a successful in-house data analytics platform. His career spans academic, commercial and philanthropic domains with a focus on innovation and delivering impactful data-driven solutions.
